Our Current Projects
click the section title in order to learn more about that particular project...
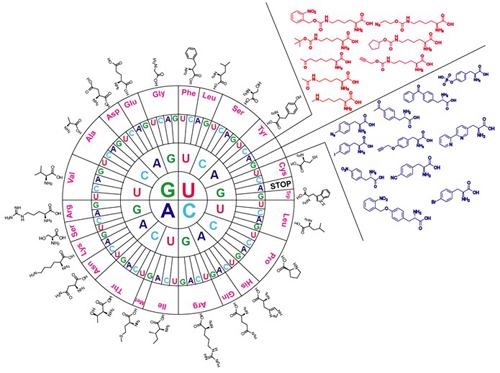
Bioorthogonal reactions refer to chemical reactions that were performed in biological system without interfering with complex cell environment. There are several catalyst based or catalyst free bioorthogonal coupling reactions have been reported. Noncanonical amino acids genetically incorporated by amber suppression encoding method can install useful functional group for the purpose of bioorthgonality in protein labelling. Several important bioorthogonal reactions have been reported dedicating in synthesis of , ex. potential click chemistry 1,3-dipolar cycloaddition between azides and cyclooctynes (also termed copper-free click chemistry) and Staudinger ligation through p-azido-L-phenylalanine (N3F) (1) genetic incorporation, oxime/hydrazone formation from aldehydes and ketones through p-acetyl-L-phenylalanine (AcF) (2) or (S)-2-amino-8-oxononanoic acid (KetoK) (3), Suzuki-Miyaura coupling and Sonogashira coupling reaction through p-iodo-L-phenylalanine (IF) (4) and Michael addition reaction by free thio molecules through dehydroalanine moiety oxidized from Se-alkylselenocysteine (SeCbzK) (5) incorporation.
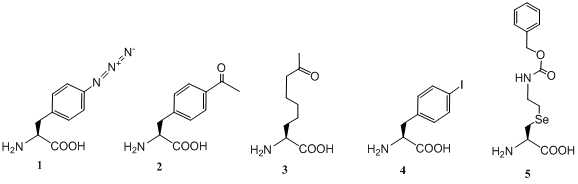
Scheme 1: Noncanonical amino acids for genetic incorporation and further chemical labelling.
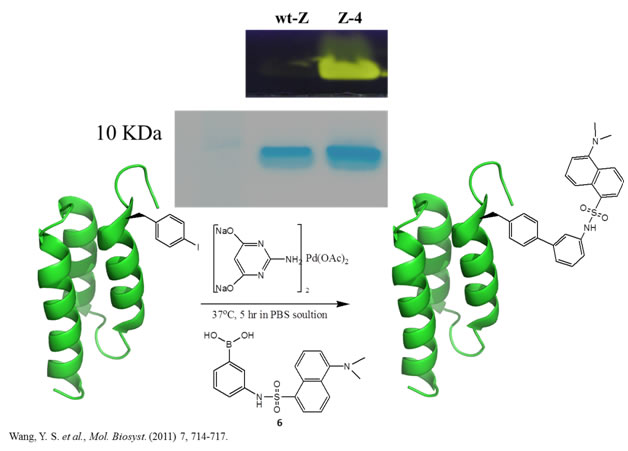
Figure 1: Site-selective labeling of Z-domain incorporated with p-iodo-l-phenylalanine (Z-4). (A) The Suzuki-Miyaura cross-coupling reaction between a p-iodo-l-phenylalanine-containing protein and 3-(dansylamino)phenylboronic acid (6). The Gelcode blue staining of Z-4 and the wild-type Z-domain protein (Z-wt) that were labeled with 6. The fluorescent imaging of the same gel when excited by 365 nm UV light. The fluorescent image shows real colors captured by a regular camera.
We are interested in understanding the genetic transcriptional control in biological system. Particularly, we have special focus on the chromatin biology. Histone unites wrap around DNA to form a compact chromatin structure. The post-translational modifications (PTMs), such as methylation, acetylation, ubiquitylation etc., on the specific residues of histone alter the chromatin motif and directly regulate the on/off of the gene transcription. This sort of study is able to provide profound information to understand cancer biology and genetic disorders and may inspire new therapeutic methods to cure these diseases.
To this specific field, we are developing methods to synthesize histone with homogeneous and defined PTMs at specific site. By means of genetic noncanonical amino acids (NAAs) incorporation, we are able to install methyllysine, and non-hydrolysable acetyllysine analogue ketolysine into protein at specific site. Besides, we are able to incorporate a precursor NAA of dehydroalanine into protein. After selective oxidation and derivatization, the PTM analogues such as monomethyllysine, dimethyllysine, trimethyllysine, acetyllysine as well as phosphoserine and phosphocystine were able to be displayed on the histone H3 at specific site. We have long-term research projects going on to study the cross-talk between different PTMs on the histone.
Phage display has contributed to determine novel and natural ligands for various targets and study protein-protein interactions, determine tumor antigens, protein–DNA interactions, in vitro protein evolution, immunotherapy and vaccine development. Phage display is highly efficient to identify peptides from large polypeptide libraries for therapeutic interventions. Development of phage display methods will make it possible to target essential components and pathways within many different diseases, including cancer, AIDS, cardiovascular disease, and autoimmune disorders. However, a phage display peptide library has limited structure motifs and functional groups because only 20 natural amino acids can be used to generate a library. We use phage and phagemid to expand the chemical diversity of a phage display library by incorporating multiple noncanonical amino acids and chemically modifying them to extend functional diversities. Screening the unnatural phage display library against therapeutic targets such as c-Abl tyrosine kinase is expected to identify highly potent inhibitors. We developed a general approach to the generation of polypeptide libraries containing multiple unnatural amino acid building blocks with phage display.
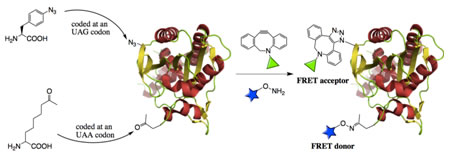
Förster resonance energy transfer (FRET) between a pair of donor and acceptor dyes is an invaluable tool to study dynamic protein conformational changes such as conformation rearrangement and folding/unfolding. The efficiency of energy transfer that is dependent on the distance between the two dyes not only represents the conformational distributions but also reflects their change upon time. Two methods are usually applied to achieve dual labeling of a protein with a FRET pair. One is to genetically fuse two green fluorescent protein (GFP) variants at the N- and C-termini of a protein. The other is to modify two cysteine residues in a protein with a small-molecule FRET pair. The GFP labeling approach suffers limitations such as labeling restriction at two termini and large sizes of GFP molecules. The cysteine labeling requires mutating all non-targeted cysteine residues, and is therefore not applicable for proteins in which cysteine residues serve critical structural and/or functional roles. In order to develop a protein labeling approach that does not have issues related with the existing methods, we plan to genetically incorporate two different noncanonical amino acids into one protein at one amber mutation site and one ochre mutation site followed by selective labeling of the two noncanonical amino acids to form a FRET pair in the protein. This method will be further used to develop biosensors for small molecules and enzymes.